Page 1, 2
PopUp Window Showing Data for Alpha Lipoic Acid
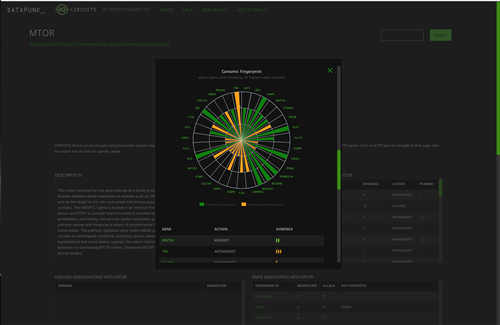
Test-Driving Circuits
Readers are encouraged to 'surf' Circuits and explore the target genes that seem more interesting. Click away! However here are a few hard links to help get you started.
- The ABO 'secretor' gene (FUT2)
https://www.datapunk.net/circuits/index.pl?FUT2
- Mitochondrial enzymes that catalyze the oxidative deamination of amines, such as dopamine, norepinephrine, and serotonin (MAOA and MAOB)
https://www.datapunk.net/circuits/index.pl?MAOA
https://www.datapunk.net/circuits/index.pl?MAOB
- Catechol-O-methyltransferase (COMT) catalyzes the transfer of a methyl group from S-adenosylmethionine to catecholamines, including the neurotransmitters dopamine, epinephrine, and norepinephrine.
https://www.datapunk.net/circuits/index.pl?COMT
- PPAR-gamma is a regulator of adipocyte differentiation. Additionally, PPAR-gamma has been implicated in the pathology of numerous diseases including obesity, diabetes, atherosclerosis, and cancer.
https://www.datapunk.net/circuits/index.pl?PPARG
- MTHFR catalyzes the conversion of 5,10-methylenetetrahydrofolate to 5-methyl-tetra-hydrofolate, a co-substrate for homocysteine (HCy) remethylation to methionine.
https://www.datapunk.net/circuits/index.pl?MTHFR
T he Code he Code
The server-side portion of Circuits was written in the Perl language, the 'Swiss Army Chainsaw' of bioinformatics. Client-side elements, such as network depictions and graphic displays of information, were coded in JavaScript using the Cytoscape JS and HighCharts JS frameworks. The PPI network was normalized using the Graphviz graphing package and the CPAN Graph module.
The Easter Egg
Users can store particularly interesting or important genes in a non-tracking cookie 'wallet,' for long-term use, thus allowing them to retrace prior investigations.
I hope the Townsend Letter readers have half as much fun exploring Circuits as I did envisioning and coding it. We at the Pathfinder Scholar Program at the COEGM are planning on expanding our open-source offerings to include a microbiota explorer that uses taxon interaction networks and Markov chains to produce a multigenerational approach to eubiosis; and a small metabolite (metabolome) explorer that employs machine learning classifiers to generate metabolic patterning characteristics. I'll make sure to alert the readers when these tools become available.
Circuits: Dataset Linkages
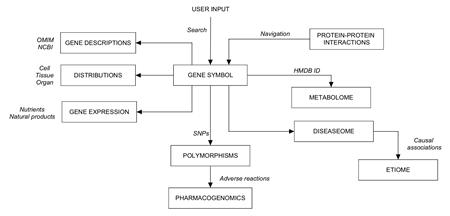
Page 1, 2
References
1. Wimsatt W. Re-Engineering Philosophy for Limited Beings: Piecewise Approximations to Reality. Cambridge: Harvard University Press. 2007
2. Landrum MJ, et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018 Jan 4. PubMed PMID: 29165669.
3. Prasad, TSK, et al. Human Protein Reference Database - 2009 Update. Nucleic Acids Research. 2009;37, D767-72.
4. Liu YI1, Wise PH, Butte AJ. The "etiome": identification and clustering of human disease etiological factors. BMC Bioinformatics. 2009 Feb 5;10 Suppl 2:S14.
5. Kaplun A, et al. PGMD: a comprehensive manually curated pharmacogenomic database. The Pharmacogenomics Journal. 2016;16:124–128.
6. Thul PJ, Lindskog C. The human protein atlas: A spatial map of the human proteome. Protein Sci. 2018 Jan;27(1):233-244.
7. Barabási A-L, Gulbahce N, Loscalzo J. Network Medicine: A Network-based Approach to Human Disease. BMC Bioinformatics. 2009; 10(Suppl 2): S14.
8. Wishart DS, et al., HMDB 4.0 — The Human Metabolome Database for 2018. Nucleic Acids Res. 2018. Jan 4;46(D1):D608-17. 29140435
Peter J. D'Adamo, ND
Distinguished Professor of Clinical Medicine and Bioinformatics
Center of Excellence in Generative Medicine
University of Bridgeport, College of Naturopathic Medicine https://www.coegm.com/ |
![]()
![]()
![]()
![]()








